The virtual laboratories enhance learning experiences by providing the student with a supplement to the physical lab. The laboratories allow students to perform exercises as in an actual lab and to gather data for preparing lab reports. To increase student's engagement and interest, they are allowed to make errors and take wrong directions, and then backtrack to correctly perform the exercise. The architecture of the system is modular and can be easily extended to implement different laboratories and it supports augmentation by animation effects and realistic rendering of virtual objects. Most of the system components, including all the virtual labs, are implemented as Java Beans. The software framework is lightweight and can be downloaded as an applet in a browser. Extensible Markup Language (XML) is used for application and data description; students can save their lab reports in XML and review them later. To demonstrate the potential of the architecture, we have developed several virtual laboratories for cell division, centrifugation, spectrophotometry, and virtual microscopy. We report experience in developing and deploying such virtual laboratories as well as their actual usage in the Department of Cell Biology and Neuroscience at Rutgers University.
Innovations in multimedia and computer-related technology offer exciting opportunities to improve the quality of teaching and learning science for students. Colleges and universities across the world are faced with a generation of students flocking to learn more about the life sciences. Consequently, instructors are confronted with more students and larger classes, particularly at introductory levels. Here at Rutgers, the General Biology course services a large group of students (in excess of 2000) with varied backgrounds and abilities. The faculty challenge is to motivate and excite these students so that each performs to their fullest potential. We believe that this seemingly difficult objective can be best achieved by incorporating computer-related, web-based, and multimedia technology into the classroom and laboratory environment.
The fundamental guideline under-guiding the development and organization of virtual laboratories is that students learn science best by experimenting and gaining hands-on experience¾raising questions, solving problems and finding answers to questions through designing and carrying out experiments. Our long-range objective is to develop virtual learning experiences for students by allowing them to access, visualize, and manipulate instrumentation and multimedia data utilizing the newly available high-speed networks. We see the development of virtual laboratory exercises as key to raising student interest and improving learning. When done effectively, this increases student access to knowledge and enhances performance as well as the quality of educational outcome.
A virtual laboratory is a virtual reality environment that simulates the real world for the purpose of discovery learning. A flight simulator is an example of a virtual laboratory. It provides a pilot with useful virtual experience based upon flying a specific type of aircraft. Virtual labs can be used in a similar fashion to augment real laboratory experience in science and technology. However, programming virtual laboratories is a very tedious and costly task. A central challenge is to develop tools allowing for rapid and easy creation of virtual laboratories. This paper presents a software architecture for rapid development of virtual laboratories.
The paper is organized as follows. Section 2 reviews background and related work. Next, Section 3 presents the software architecture of the framework. Section 4 gives examples of several virtual biology laboratories implemented using the framework. Section 5 presents some observations and results obtained from students who actually used the virtual labs. Finally, Section 6 concludes the paper.
The need for multimedia technology in biology teaching is being recognized worldwide. There are several interactive virtual labs currently available on the web. California State University's Center for Distributed Learning [2] now has eight biology labs in the beta test phase on topics ranging from evolution to molecular biology. The labs are commercially marketed through the Addison Wesley Longman publishing house. A series of interactive Java tutorials are offered at [4] that explore various aspects of virtual Scanning Electron Microscopy (vSEM). The student can discover how specimens appear when magnified in the virtual SEM. Each time a new specimen is loaded into the browser, the focus, brightness, and contrast controls are randomly reset to simulate the situation in a real microscope. The University of Melbourne has established a Science Media Teaching Unit [15] to promote effective and efficient use of computer-based multimedia in teaching biology. Physics 2000 [8] comprises interactive Java applets through which students can explore elementary physical phenomena. Edmark [6] markets interactive virtual labs that are designed to make it easy for teachers to integrate the programs into their curriculum. Unlike our labs, it appears that these are not platform independent Java applets. Hence a student would have to download a version specific to the platform she is using. Researchers at the TeleLearning Network of Centres of Excellence [5] are developing virtual labs that enable students to carry out a range of experiments by computer, thus enhancing their classroom learning. Here, students use a mouse to select objects in the lab, move them around, and adjust parameters such as the intensity of an electrical current or the frequency of a laser. As the instruments in the actual lab can be rather fragile and expensive, the virtual labs are a reasonable alternative solution for enabling the students to "play" with the instruments. Just like their real equivalents, the virtual instruments respond to students' manipulations by providing correct data if the experiments have been carried out properly. These labs are implemented as Java applets but there is no account of a generic underlying software architecture that simplifies the lab development. The researchers are also pursuing the development of remote labs, where robots receive commands from students and reproduce the virtual experiments in a fully equipped real lab. Perhaps the most advanced biology lab exercises are offered through the Howard Hughes Medical Institute. They developed Bio-Interactive [9]¾a collection of learning modules that let students interactively explore topics in cardiology, neurophysiology and immune system. These virtual exercises augment neuroscience laboratories and have been received with enthusiasm by faculty and students. In these exercises, user-action is restricted mostly to "clicking" on the various instruments. For a better realistic feel, the users should be able to perform the exact operations as they would on the real instrument.
We notice in most of the aforementioned work the widespread use of slider widgets. In the actual instrument, this need not be the case. For instance, we might need to "rotate" a dial to set a particular reading rather than "slide" a marker across a slider. We attempt to overcome these shortcomings and provide the user with a more realistic experience through the virtual lab. Also, although the above reviewed exercises are "interactive," they are very linear. Our goal is to develop exercises that give student choices and options such that two students may not have the "exact" same lab experience, but finish having learned the same concepts.
The architecture is based on the Java Beans framework [14] and extends our prior work on using Java Beans in collaborative applications [11]. The class of beans that follow this architecture is called Manifold [12].
The main characteristic of the Manifold beans is the multi-tier architecture [12]. The common three-tier architecture comprises the vertical tiers of presentation, application or domain logic, and storage. The Manifold's presentation tier is virtually free of the application logic and deals with visualizing the domain data and accepting the user inputs. The domain tier deals with the semantics of tasks and rules as well as abstract data representation.
The domain tier contains all the model objects and the presentation tier contains all the view/controller objects of the MVC design pattern [1]. The main benefit of this decomposition is the resulting separation of concerns. The design internally consists of three beans, although the outside world sees a single bean.
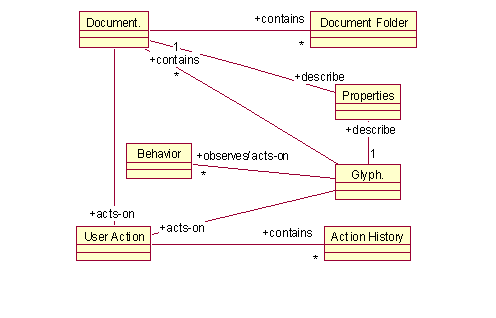
The conceptual model of a typical document editor is shown in Figure 1. The key concept is a Glyph, which represents all objects that have a geometry and may be drawn. The name "glyph" is borrowed from typography to denote simple, lightweight objects with an instance-specific appearance [7]. A Glyph is essentially a container for a list of <property, value> pairs, with properties such as dimensions, color, texture, etc., but also various constraints on glyph manipulation. A Glyph may be an aggregate containing multiple leaf or aggregate glyphs thus embodying the Composite pattern [7]. A corresponding (simplified) class diagram is shown in Figure 2.

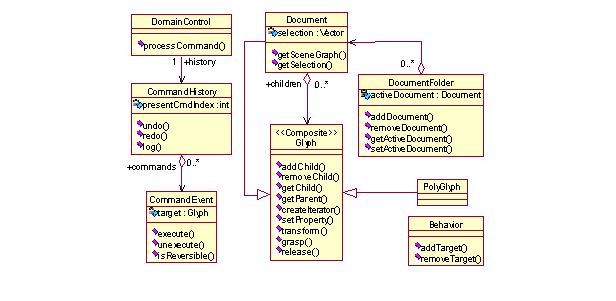
All Glyphs in a document form the scene graph, itself a Glyph, which has a tree data structure. The scene graph is populated with different vertices (Glyphs) in specific applications that extend the Manifold framework. Glyphs are divided into two groups. Leaf Glyphs (terminal nodes) represent individual graphic elements, such as images, geometric figures, text or formulas in spreadsheet cells. PolyGlyphs are containers for collections of Glyphs. They correspond to branch nodes and can have children. Example PolyGlyphs are group figures, paragraphs, maps or calendars. PolyGlyphs have all the functionality of Glyphs. They also have the additional property that they can contain Glyphs or other PolyGlyphs. For example, in the virtual spectrophotometer lab (described in Section 4 below), an example of PolyGlyph is a control dial, which contains an ellipse figure (knob) and a line figure (reference mark on the dial). Another example, a map PolyGlyph positions an icon Glyph according to its (x, y) properties, whereas a card pack PolyGlyph positions all its Glyphs stacked one on another, disregarding the Glyph's coordinate properties.
Glyphs are sources of the following types of events that are fired in response to the operations on the scene graph tree structure: AppearanceEvent for Glyph add/remove operations, PropertyChangeEvent for changing the Glyph properties, and TransformEvent for applying the affine transforms on the Glyphs. The interested parties register as event listeners for some or all types of the events via the Java Beans delegation event model.
DomainControl is the system controller for the domain bean that invokes the system operations. This is the only portal into the domain bean. The only way to cause a state change in the bean is to invoke the processCommand() method. Even the local presentation (view) objects interact with the domain objects through this portal only.
The CommandEvent class implements the Command pattern [7] and has the responsibility of keeping track of the argument values to invoke operations on Glyph and Document objects so the operations can be undone/redone. We name this class CommandEvent instead of Command to emphasize the Java event distribution mechanism. Commands create/delete Glyphs or correspond to Glyph methods. In addition, we have commands to open or save a document and document-view-related commands.
Behaviors are objects that observe the Glyphs as event listeners and act on Glyphs by invoking the processCommand() method on the DomainControl. Behaviors maintain a list of named target Glyphs that are acted upon. Example behaviors are collision detection in three-dimensional worlds, spreadsheet cells with formulas, or coordinated manipulation of several Glyphs, which is not the same as a group movement where all objects are manipulated in the same manner. Unlike the Java 3D behaviors [13], which are oriented towards avoiding the unnecessary rendering of invisible parts of the world, our behaviors are focused on end-user programmability. The user "wires" the behavior to the event sources and the targets as will be seen in Section 3.3 below.
The Model-View-Controller (MVC) design pattern divides an interactive application into three components [1]. The model contains the core functionality and data, views display information to the user, and controllers handle user input.
A Glyph may have a corresponding GlyphView, which is a view part of the MVC pattern associated with the model Glyph. The reverse may not be true, depending on whether or not the containing Document notifies its AppearanceListeners (DocumentView) about the creation of a new Glyph. The GlyphView subscribes to the model and listens to the important state changes. Thus, the derivatives of this class may implement some or all of the listener interfaces (AppearanceListener, PropertyChangeListener, and/or TransformListener) as needed.
The key user activity in graphical user interfaces is direct manipulation of screen objects. The classes that support direct manipulation are Tool and Manipulator [16], Figure 3. Tool encapsulates information about the current direct manipulation mode, e.g., rotation, resizing, etc. Tools are essentially state objects for DocumentViews (see the State pattern in [7]). Manipulator encapsulates a Tool's manipulation behavior and is responsible for providing visual feedback during a manipulation sequence (e.g., redrawing a rubber-band using the XOR technique). The Tool-Manipulator breakup separates the state information from manipulation behavior. Manipulation involves a sequence of grasp-wield-effect operations, each of which results in a message to the manipulated object, which is encapsulated in a CommandEvent.
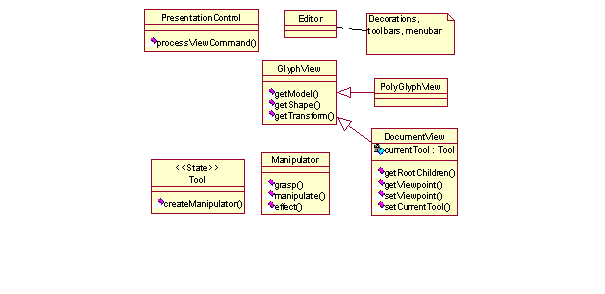
Manipulator is the Controller part of the Model-View-Controller design pattern in that it converts the user interaction into the CommandEvents for the model. PresentationControl gathers all user actions originating in the presentation bean as CommandEvents and delivers them to its CommandListener(s), normally a DomainControl object. PresentationControl is the system controller that processes the presentation-related CommandEvents, such as changing the viewpoint, that originate at a remote process. Manipulator separation helps keeping the application lightweight (especially presentation layer), since the Manipulators are created only for direct manipulation.
In order to provide for end-user customization of the application, we need to specify a data and application description language. The language is used to describe the data being operated upon, i.e., the initial scene graph of the application, as well as the relationships between the application objects. The language should be rich, yet easy to use and fast to parse. Since the scene graph is a hierarchical structure, the language should accommodate hierarchical data. The World Wide Web is at present the predominant means of exchanging information and delivering documents between networked domains. XML (eXtensible Markup Language) is now being promoted as a new Web markup language for information representation and exchange [17]. It satisfies all of the above listed requirements and also has been used for application description [10]. Thus, our choice for data and application description language is XML.
An end user or an XML programmer creates Manifold XML files based on the set of available Glyphs and their attributes. The correspondence between the elements and Glyphs is not one-to-one. Not all XML elements are Glyphs. For example, Glyph properties may be represented as sub-elements of the Glyph element. Here is an example Glyph EllipseFigure from a two-dimensional graphics editor, called Flatscape, that is based on the Manifold framework:
|
<GLYPH type="flatscape.domain.EllipseFigure">
<PROPERTY name="glyph.height" type="java.lang.Double" value="42.0" /> <PROPERTY name="glyph.width" type="java.lang.Double" value="42.0" /> <PROPERTY name="fill.color" type="java.awt.Color" value="java.awt.Color[r=150,g=150,b=150]" /> <TRANSFORMATION type="flatscape.domain.Transform2D" value="0.0 0.0 1.0 1.0 0.0" /> </GLYPH> |
The sub-elements could even have their own sub-elements if, e.g., the transformation is represented as independent scale, rotation and translation parameters, each tagged individually. In addition to Glyphs, the user can specify the Behaviors. Each behavior may listen to Glyphs for events (AppearanceEvent, PropertyChangeEvent, and TransformEvent) and may have specified targets onto which it acts (other Behaviors or Glyphs). Here is an example:
|
<BEHAVIOR id="steering" type="bicycles.domain.Steering">
<LISTENER type="manifold.domain.TransformListener" source="handlebars" /> <TARGET name="wheel" ref="frontWheel" /> </BEHAVIOR> |
The Behavior object labeled "steering" listens to the Glyph labeled "handlebars" for TransformEvents and acts on the target labeled "frontWheel." As the user manipulates the handlebars, the behavior receives the transform events, computes the rotation angle for the front wheel of the bicycle and sends a TransformCommand to the wheel. The behavior classes, such as bicycles.domain.Steering in the above example, are the pre-existing Java classes or must be programmed by the end-user in Java.
A key benefit of implementing presentation and domain as distinct beans rather than the whole package as a single bean is in being able to mix and match different combinations. We can have a set of more or less complex beans for each. Different domain beans can implement complex behaviors and the presentation beans can implement visualizations with varying realism.
The virtual lab contains a set of objects such as microscopes, centrifuges, whole organisms, or individual cells each with specific pre-programmed behaviors. The student interacts with the objects in order to attain a set of given goals, i.e., study of cell features, separation of cellular components, measurement of enzyme activities, quantification of cell division, etc. The use of creative renderings of objects and their behaviors allows the student to freely experiment in the virtual world. Module content of virtual labs, complexity of problem solving, and sophistication of technical skills are vertically scaled so that each student can move through the module depending upon level of preparation (from General Biology student to advanced students in Fundamentals and Advanced Cell Biology).
The Manifold framework significantly simplifies the development of virtual biology laboratories. The developer's main task is in writing the XML document and programming the Behaviors associated with the lab. Currently we have implemented five virtual laboratories and the amount of lab-specific Java code relative to the Manifold code ranges from 5% to about 20% in very complex labs. In addition, the new code is highly standardized, relieving the developer from the issues with display, document parsing, etc., and only requiring the developer to program the particular Behavior classes.
We also make use of the CommandHistory facility to provide the student with a Back button by which he/she can backtrack and perform the previous actions again. Each lab stage is marked and all the events that occur between two stages are logged. When the student clicks on the Back button, all the events that occurred after the last stage are undone, of course, if these events are reversible. Students can use a powerful graphics editor available in the framework to prepare lab reports after the exercises. Any stage of the lab can be captured and copied in the report document at the level of structured graphics, rather than screen bitmaps. The documents are stored in XML and can be reviewed and edited manually if necessary. The following sections review some of the currently implemented virtual laboratories.
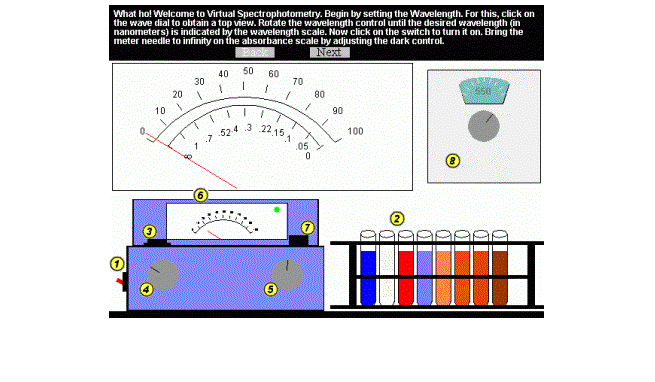
The spectrophotometer is a machine used to measure the concentration of a substance in a solution by passing light of a specified wavelength through it. A photocell at one end receives the transmitted light and an analog meter displays the percent transmittance of light received by the photocell. In order to use this machine, the correct wavelength of light must be chosen so that the light is absorbed by substance when it tries to pass through the solution (the more substance there is, the less light will pass through and vice versa). The main purpose of the lab is to familiarize students with the spectrophotometer and its use. The students first calibrate the instrument using a reference solution of known concentration. The concentration of unknown samples is calculated according to the transmittance value of this solution. Here is an example of how the lab developer specifies the zero control dial (Figure 4) as a PolyGlyph consisting of a circle and a line, which represent the knob and the reference mark:
|
<POLYGLYPH id="zeroDial" type="flatscape.domain.PolyGlyph2D">
<PROPERTY name="glyph.permittedUserTransform" type="java.lang.String" value="rotate" /> <PROPERTY name="glyph.dialType" type="java.lang.String" value="zeroControl"/> <TRANSFORMATION type="flatscape.domain.Transform2D" value="79.0 495.0 1.0 1.0 0.0 0.0 6.5" /> <GLYPH id="zeroDialKnob"
type="flatscape.domain.EllipseFigure">
<GLYPH id="zeroDialReferenceMark"
type="flatscape.domain.LineFigure" >
|
The remaining objects are specified in a similar manner. The developer must also specify the behaviors, as in this example:
|
<BEHAVIOR id="measuring" type="biology.spectro.domain.SpectroMeasure">
<TARGET name="pilotLamp" ref="pilotLamp" /> <TARGET name="needle" ref="needle" /> </BEHAVIOR> <BEHAVIOR id="turning" type="biology.spectro.domain.DialWatcher">
<BEHAVIOR id="opening" type="biology.spectro.domain.LidWatcher">
|
The Behavior "measuring" performs the calculations based on the solution's density and the light wavelength and sends a TransfromCommand to the instrument (AbsorbanceMeter) needle to display the wavelength. It also turns on or off the pilot lamp when the spectrophotometer is turned on or off. The Behavior "turning" causes the Behavior "measuring" to redo the measurement when a dial is rotated. Similarly, the Behavior "opening" causes the Behavior "measuring" to redo the measurement when the sample holder's lid is opened or closed.
In the actual lab, students are given plastic beads and a cylindrically shaped magnet called a "centromere." When these parts are assembled, they form the model for a chromosome. The students are asked to build four of these, one red pair, and one yellow pair and explore different phases of cell mitosis and meiosis. In each of these phases, the chromosomes behave in a certain way in the cell while the cell first divides into two and then into four cells. The students are asked to manipulate the chromosomes to show their behavior during each of these stages. Screen captures in Figure 5 show selected phases of the meiosis virtual lab. Any time the cell components are arranged in a particular configuration, the behaviors are set in motion and perform animation of the corresponding cell process.
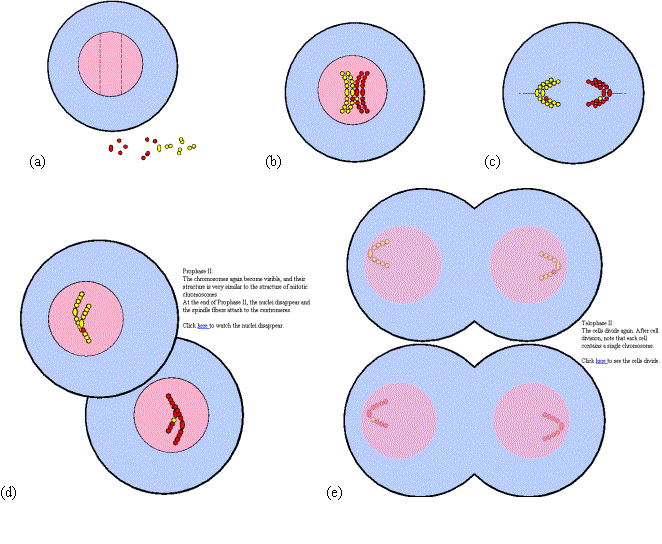
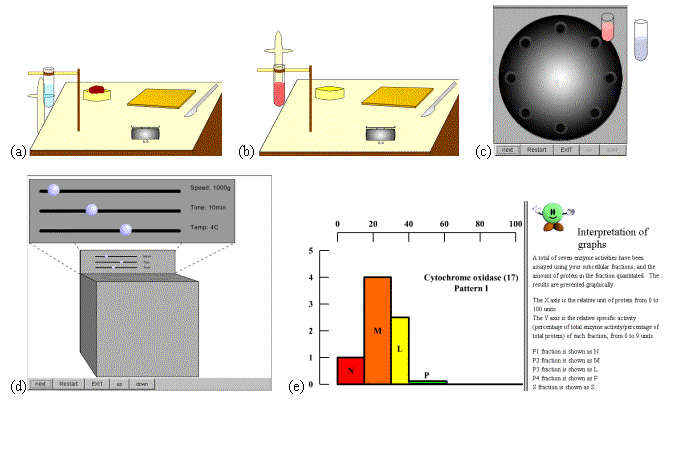
Differential centrifugation is a mode of centrifugation in which the sample is separated into two fractions: (1) a pellet consisting of sedimented material and (2) a supernatant. The experiment is similar to the one originally described in 1955 by Christian de Duve for the discovery of the organelle lysosome (awarded a Nobel prize in 1974). It is based on the differences in sedimentation rate of particles of different size and density. The tissue homogenate is centrifugally divided into a number of fractions by stepwise increasing the applied centrifugal field. The centrifugal field is chosen so that a particular type of organelle will be sedimented as a pellet, and the supernatant will be centrifuged at a higher centrifugal field for further fractionation. Our virtual lab simulates the operation through the following major stages (Figure 6):
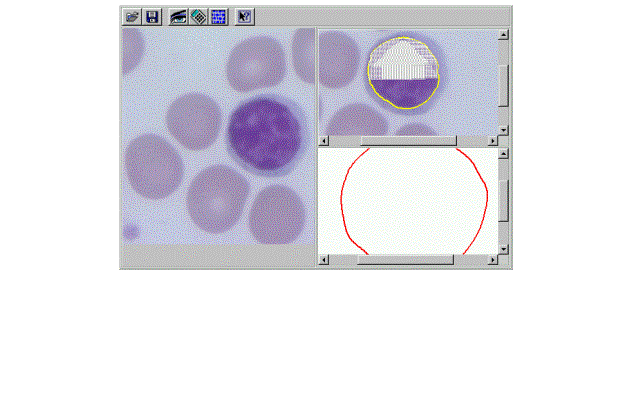
This laboratory was developed in collaboration with the University of Medicine and Dentistry of New Jersey. The lab allows the student to load an image and view it at different magnifications (Figure 7). The left panel shows an original image of blood cells, in this case afflicted by leukemia. Automatic techniques for image segmentation are called up by pen-based gesturing and by speaking voice commands. Image analysis methods¾developed in the Robust Image Understanding Laboratory at the CAIP Center [3]¾can extract common components on the basis of color and texture (the top small panel), and by edge shape (the lower small panel). We are currently working on connecting the lab with the physical microscope for real-time image acquisition. The controls will be provided to manipulate the microscope remotely.
The core biology course in the Department of Cell Biology and Neuroscience at Rutgers University has been a testbed for the cell mitosis lab over the last semester and will continue to be so, additionally using the new labs (cell meiosis, differential centrifugation, spectrophotometer, and a virtual microscope), which have recently become available. The short-term goal of virtual labs is to serve as a preparation and supplement for actual labs. The students are thereby familiarized with procedures before they actually go into the lab and perform experiments. This "rehearsal" by simulation of a complex experiment is a cost-effective preparation for the use of limited and expensive lab facilities. The laboratories are continuously available on our web site so the students can access the labs from anywhere. Our labs also enable self-paced learning for each student. We do not keep track of how many students access the laboratories or the amount of time they use it. As these labs are available on the web anytime, students might open the labs and simultaneously refer to their textbooks in order to understand the concepts better. Hence the time for which they use the lab does not really provide any valuable information.
We present a summary of preliminary results from the ongoing evaluation of the Virtual Biology Labs by the Rutgers Department of Education at Rutgers University. Following are some trends that we are observing in student reactions to the virtual labs and how those reactions influence the lab design.
In general, students had a positive attitude towards the lab on mitosis. Of the 18 students who were surveyed, 15 commented on the usefulness of the simulations in explaining the different stages of mitosis via the dynamic representations and simulations that were embedded in this lab. Specifically, students liked the fact that they could replay and watch the process as many times as they needed. Also, the exercises and feedback provided were considered a positive feature. In reflecting on the usefulness of the spectrophotometer simulation, the students enjoyed the individual and repetitive practice they could engage in. They envisioned that this experience would help them with their lab practical exams as well as fine-tune their skills in operating the spectrophotometer in the actual lab. Overall, the students found the virtual labs to be an interface where they could learn and practice in spite of making errors. They also acknowledged that the simulations were realistic and helped in demonstrating certain processes that were not easily represented in the actual lab.
Although the feedback was generally positive, there were some features that students had trouble with. Several students expressed the concern that the instructions accompanying certain simulation tasks were unclear. In order to test the usefulness of the Back button, we have implemented it only for some labs. In cases where the Back button was not implemented, students found that in the event they made an error they were required to start the process all over again, which was frustrating. They felt that if the exercise were designed to allow one-step backtracking, it would be more accessible. This validates our assumption that the Back button is a feature that could help enhance the learning experience for the student.
Whenever we design the user interface for a virtual lab, we make certain assumptions about the user. What could be very obvious to the designer need not be intuitive at all to the end-user. Hence we performed a usability study for the virtual labs. Table 1 captures the results of a study performed for the virtual spectrophotometer lab. We have also given the possible explanations for students feeling the way they did when they used the lab. Many of the findings serve as development hints as to how to improve the design. The findings about the assumptions 11-14 in particular emphasize the need for an expert system based automatic help and guidance. Table 2 discusses the ease of use of each feature of the lab. As the students perform the physical lab as well as the virtual lab, our virtual lab should provide features that the real lab does not. A main feature in our labs is their "non-linear" nature. When students perform these virtual labs, they are likely to make errors that they might notice only in the subsequent steps. Thus, in order to give an opportunity to the user to correct the mistakes, we provide him/her with a Back button. As expected, this feature is quite helpful to the student. Table 3 tabulates the features of the virtual lab that are significantly different from the real lab as recognized by the students. As the design of virtual labs is part of on-going research, we also asked the students for feedback on the problems that they faced while using them. This will provide us with valuable insight as to how to improve our future designs. Table 4 presents some of the user comments. Table 5 presents a few informal observations made by the interface designer.
| Assumption | Valid | Invalid | Comments |
| 1. Students will know which dial is the wave dial. | 80% | 20% | As there are 3 dials, there is a possibility of confusion; again, crying out for tool tips. |
| 2. Students will know where the switch is located. | 67% | 33% | Probably because the switch was located in a different place from the actual instrument where it is combined with the zero control dial. |
| 3. Students will know which is the transparent solution. | 100% | 0 | Probably because all the other test-tubes had rather distinct colors. |
| 4. Students will know that they have to set the wavelength before calibration. | 100% | 0 | This is a little surprising because some students did not know where the wave dial was! Perhaps their answer is not true? |
| 5. Students will know how to obtain the top view. | 73% | 13% | Two students claimed that they do not remember; perhaps they did not use it at all? |
| 6. Students will know how to make the top view disappear. | 60% | 33% | Again one did not try it; here is it safe to assume that more students actually did not try to make it disappear because it did not cause hindrance? |
| 7. Students will use the top view at all | 93% | 7% | Students did try to set the wavelength. |
| 8. Students will know that they have switched on the spectrophotometer. | 100% | 0 | Students do notice the pilot lamp being turned on. |
| 9. Students will know that if they switch off, they cannot use the spectrophotometer. | 100% | 0 | Most students did try to operate the spectrophotometer without switching it on; as they saw that nothing was happening, they read the instructions to see that it has to be turned on first. |
| 10. Students will differentiate between the transmittance and absorbance scales. | 73% | 27% | As this questionnaire was provided after the students finished the experiment, students might not remember which scale they used--top or bottom. |
| 11. Students will recognize the right reading on the right scale. | 86% | 14% | As one scale is a logarithmic scale, it is the only one with an "infinity" reading; so this should be intuitive. |
| 12. Students will remember the effect of sample holder not being closed. | 57% | 43% | Though they know that the reading is wrong, they are not sure how different it will be (higher or lower reading). |
|
13. Students will notice the Back button getting automatically activated. |
60% | 27% | Two students never used the Back button because they calibrated exactly the first time. |
| 14. Students will click on the Back button when they are instructed to. | 80% | 7% | Same as above. |
| Property | 1 | 2 | 3 | 4 | 5 | 6 | 7 | Comments |
| I found the dials easy to rotate | 1 | 2 | 3 | 4 | 4 | 0 | 1 | Need a tool tip that tells the students where to click on the dial and rotate it. |
| I found the Back button useful | 2 | 1 | 3 | 2 | 1 | 2 | 4 | Some students never used the Back button; also, students wanted to backtrack each stage rather than just the single stage back. |
| I found it easy to read from the meter scales | 6 | 3 | 2 | 1 | 2 | 0 | 1 | Perhaps if we compared the student's readings with the actual readings given by the spectrophotometer then this would be more meaningful. |
| 1. In the physical lab, concepts like mitosis seem abstract. But in the virtual lab, the idea of something like mitosis is more concrete. |
| 2. It is nice that students are allowed to see what they see in lab, at home. |
| 3. The virtual lab is concise and to the point. |
| 4. The virtual lab corrects you when you go wrong; so you can learn from mistakes. |
| 5. The virtual lab has more graphics that help you understand mitosis better, while the actual lab only uses the microscope. |
| 6. We used beads in the real lab; in the virtual lab, the click and drag method with each bead was very annoying. |
| 7. The simulation actually helps us understand what we learned in the lab better. |
| 8. The physical lab had more examples and we had to construct different mitosis scenarios, including both haploid and diploid cells. |
| 1. When I did not put the chromosome bead in the right place, there should be a Help button to tell me what went wrong. |
| 2. The dialog box pops up too many times during the simulation. If you can somehow get rid of that, it would be great. |
| 3. The two windows concept sometimes creates confusion. |
| 4. There should be a forward and Back button to skip around rather than do everything allover again as was provided in the spectro lab. |
| 5. The instructions sometimes were not clear. |
| 1. People do not go through the instructions given to them initially. They mostly try to use the applet without reading the instructions. Only when they do not understand what is going on, they start reading the instructions. Thus the applet has to be pretty much self-explanatory. |
| 2. When a student makes an error, we are faced with two options: let the student go on and add an error percentage to the readings which he will know towards the end of the experiment (the Back button will be active throughout here); or to explicitly inform the user that he/she has committed an error. |
| 3. We realize that instructions are not the most eye-catching aspect of our labs; hence we should try to use something similar to dialog boxes that is more noticeable to the user. But sometimes users do get a little frustrated if a dialog box keeps popping up for every error they commit, as was commented upon in the mitosis lab. This aspect is an interesting area of future study. |
This paper presents software architecture for rapid development of virtual laboratories. The benefits of virtual labs over actual laboratories are found in their increased portability, cost effectiveness, reduced need for teacher intervention, increased student interest and control, adaptability to various learning styles and learning rates, web ready software and self-testing. Virtual labs will satisfy a growing need for engaging interactive learning software.
The architecture presented here can be used in developing tools to support scientific laboratories that allow sharing unique or expensive instruments. An important missing component is safety and security for safe operation of an instrument coupled with user authentication, privacy, and integrity of data communication. Both of these are part of our continuing work.
Our experimental findings call for the development of an expert system based automatic help and guidance in running the laboratories and this is part of our continuing research.
The virtual labs are presently single-user labs. As our framework can support collaborative work, we are working on designing collaborative laboratories or collaboratories. Scientific collaboratories enable researchers to work together across geographic and organizational boundaries to solve complex, interdisciplinary problems and to have access to remote resources. In our virtual labs, students could collaboratively perform experiments and share and compare their results.
Further information and source code are available at:
Allan Krebs, Kevin Johns, and Abhijit Bhaware contributed significantly to the development of the virtual laboratories. Prof. Richard Triemer, chairman of the Department of Cell Biology and Neuroscience at Rutgers University, initiated the project and provided great support in all phases. The research reported here is supported in part by a grant from the New Jersey Commission on Science and Technology, the DARPA Contract No. N66001-96-C-8510 and by the Rutgers Center for Advanced Information Processing (CAIP).